Overview
The QC Report is a self-contained HTML page that you should be able to open with any common modern browser:
Chrome >= 80
Firefox >= 113
Edge >= 80
Safari >= 16.4
Opera >= 67
The QC Report collects and makes easily accessible a summary of several run metrics, grouped in sections. Each section contains a different dashboard to display summary plots. The interactive dashboards display additional details for each data point when hovering the mouse over it.
The QC Report is divided in two parts:
1. A header with the sample information, main metrics and a link to contact our support.

-
Number of Cells: cells that have been called by pixelator. -
Avg. Pixelator Filtered Reads per Cell: average number of reads filtered per cell that support molecules collapsed. -
Avg. Antibody Molecules per Cell: average number of antibody molecules per cell discovered.
2. A body and a footer with the different sections and links to our social media.
Sections
Although we show the content of the body collapsed in the figure above, the different sections contain specific information you may be interested in.
Parameters: specific parameters for the run (or all parameters if selected)Cells: cell calling and general metrics at the cell levelCell annotations: cell annotations into the major PBMC lineagesAntibodies: antibody quality metricsSequencing: sequencing quality metrics
Each metric per section has an associated info button and by hovering your mouse pointer over it, a pop-up will show a message with the description of the metric:

When one of the metrics show an unexpected behavior, it will show a small warning icon. By hovering the mouse pointer on that icon, a message pop-up on the warning will appear.

Parameters

The parameters section shows the specific parameters that have been input to each stage in the nf-core/pixelator pipeline run.
You can also activate the "Show all parameters" button that will show all the parameters that are input to each stage. When all parameters are displayed, the parameters that the user has input are marked with a pencil icon.

Cells
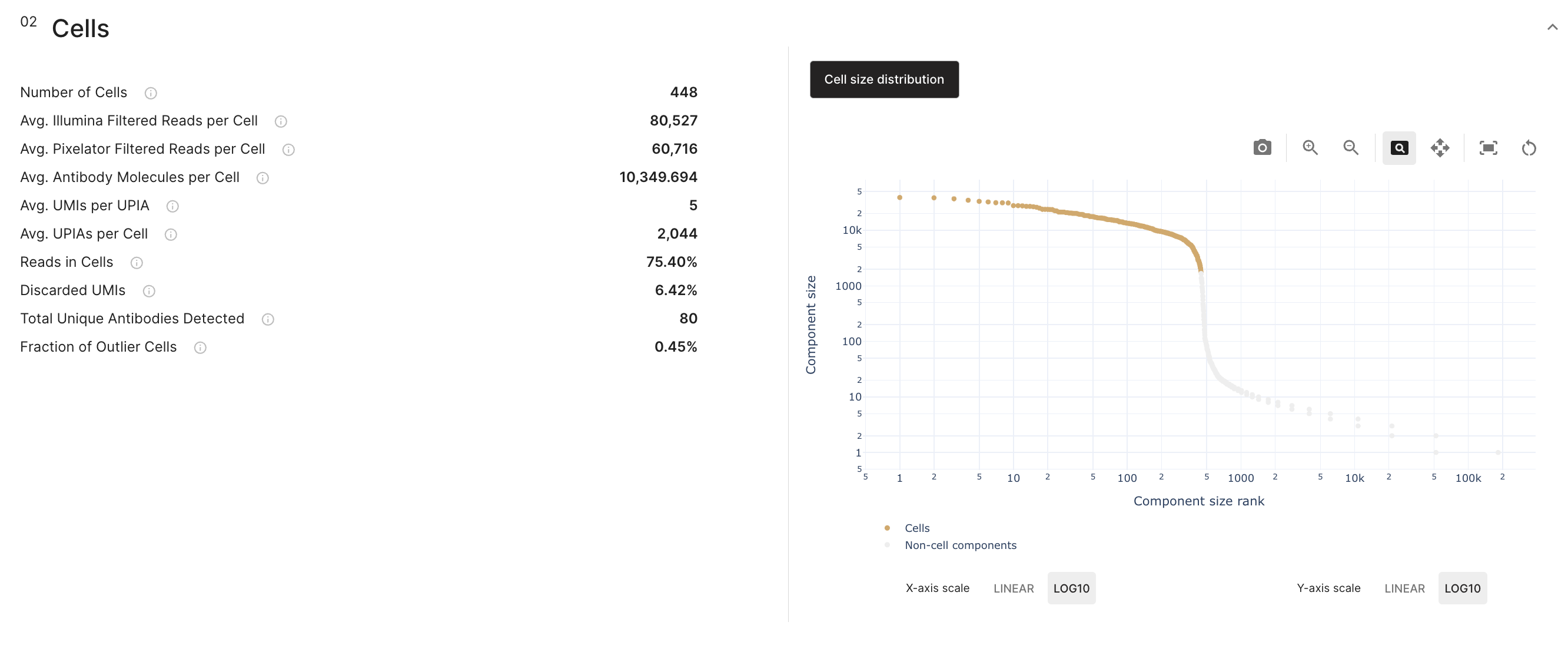
The cells section show the summary of the sample and MPX assay metrics after calling for cells.
The default view called "Cell size distribution" shows the molecule rank plot that corresponds as to where the threshold for cell calling was found. Cells are colored as a scatter plot of the decreasing distribution of their molecule sizes. The non-cell components are greyed out in the background of the distribution.
Each axis values can be displayed in a linear or logarithmic () scale. And you can switch between them.
Cell Annotations
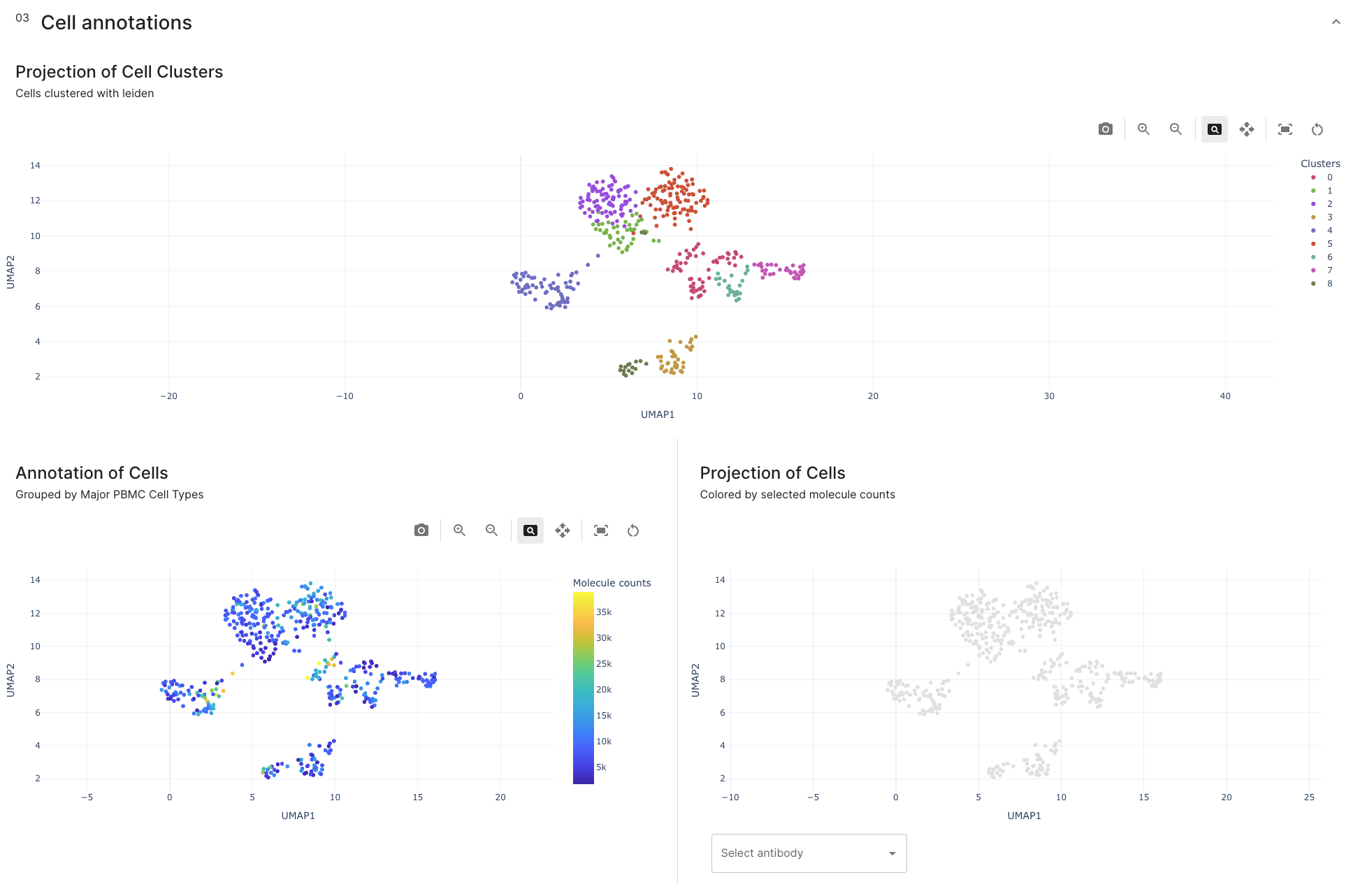
Currently three UMAP projections will give you a good overview of cell group annotations. These dashboards allow to understand better the content of your sample.
The upper UMAP projection shows the clusters created after leiden clustering. When pixelator process each sample on the annotation stage, UMAPs are created with the default scanpy parameters. Those clusters show groups of cells with similarities in terms of antibodies presence.
The lower left dashboard shows an UMAP projection of the total antibody molecule counts per cell on a color scale.
On the lower right dashboard, different major groups of PBMCs can be identified by the amount of certain antibodies present on cells.
On this dashboard, you can search for markers of interest, one at a time. There is also search functionality that allows you to search between the antibodies in the panel list.
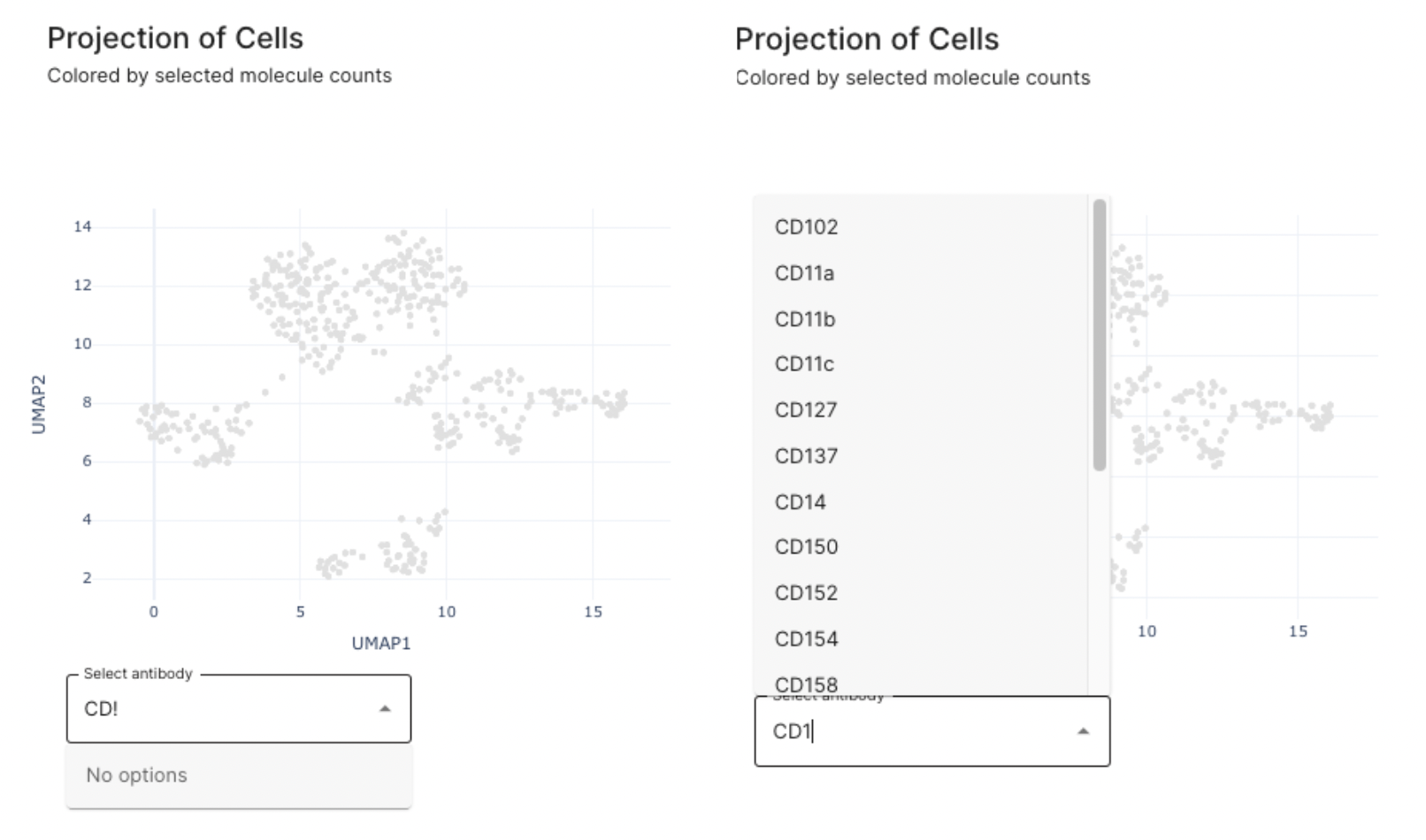
Only one antibody at a time can be selected. Both lower panels present raw molecule antibody counts.
The functionality across these UMAP panels allow MPX customers to look for certain antibody specificity and validation in grouped clusters of cells.
The bottom dashboard panel in this section represents count information per antibody in violin plots. However, this information is transformed in a scale for better comparison.
The information can be sliced across major groups of PBMCs if annotated. If no annotations are available, cells will be grouped as "Unknown cells".
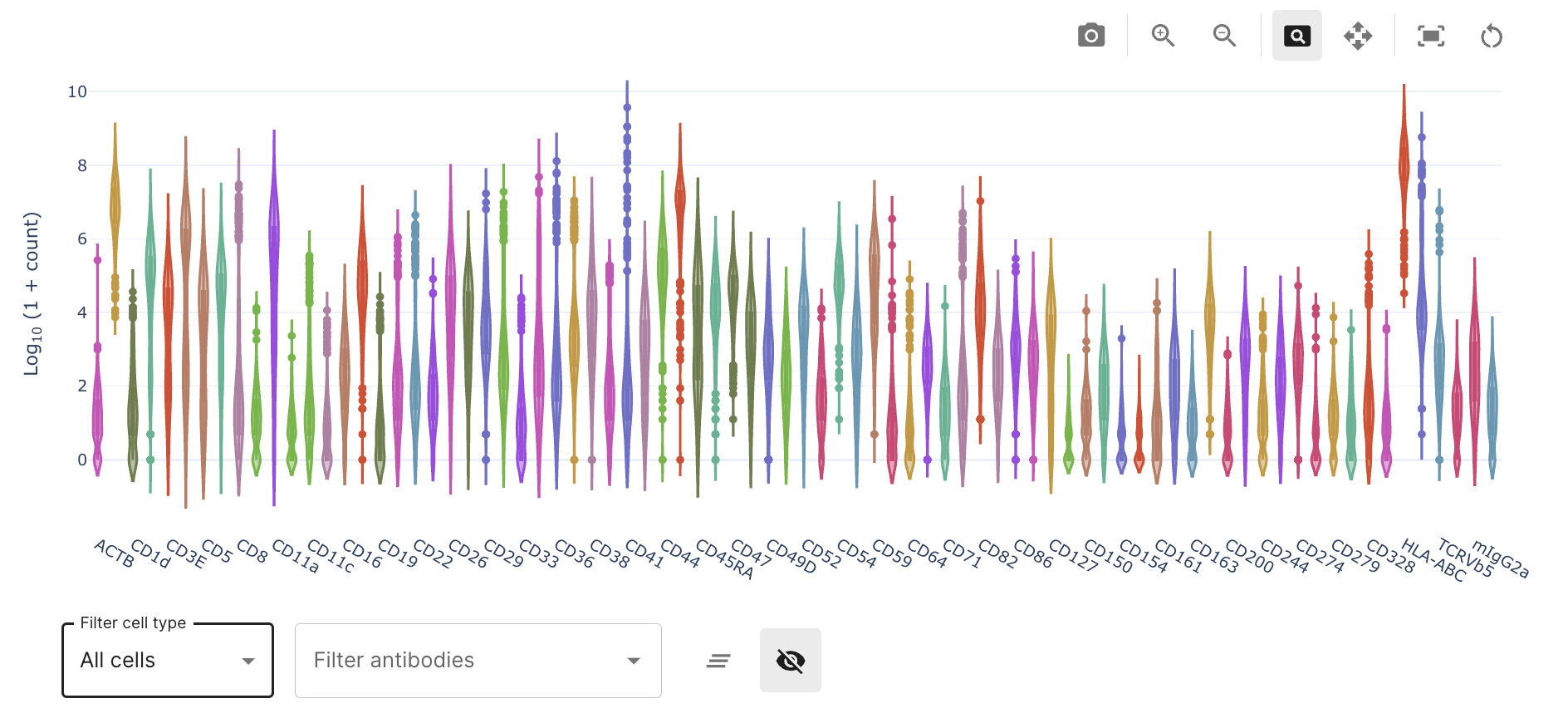
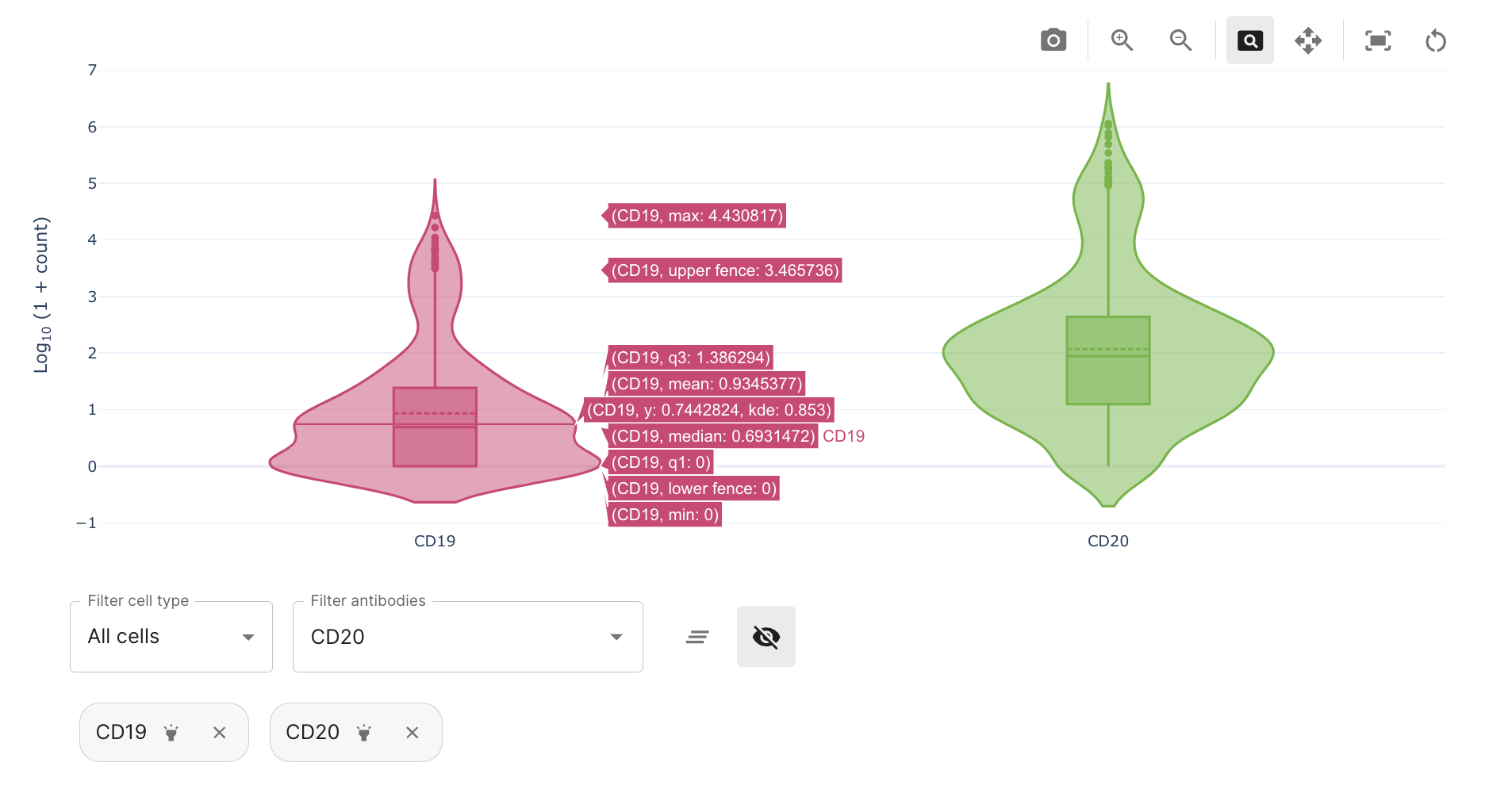
Also, several antibodies can be further selected in the "filter antibodies" menu. An active selection can be deselected if those antibodies if not important anymore. This functionality is a useful tool to study antibodies with known cell type annotations and their level of signal and background in the sample (e.g. CD19 and CD20 for B-cells). Plots in dashboards are interactive and in the case of violin plots, several summary metrics can be extracted.
Antibodies
The antibodies section describes several of the metrics of reads containing antibody information either if in finally called cells or just components.
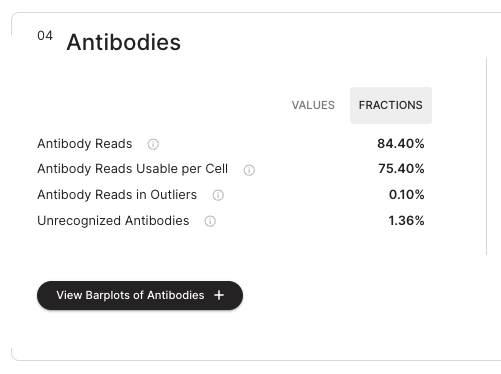
The user can toggle between percentages and absolute values of the metrics with a toggle.
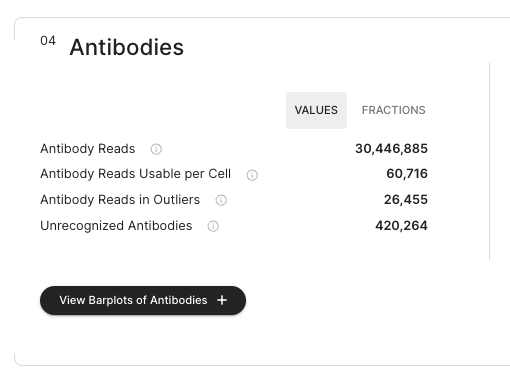
If the user expands the "View Barplots of Antibodies", the antibody section displays a dashboard with percentages of the number of antibody reads that have been decoded and assigned to each antibody identity.
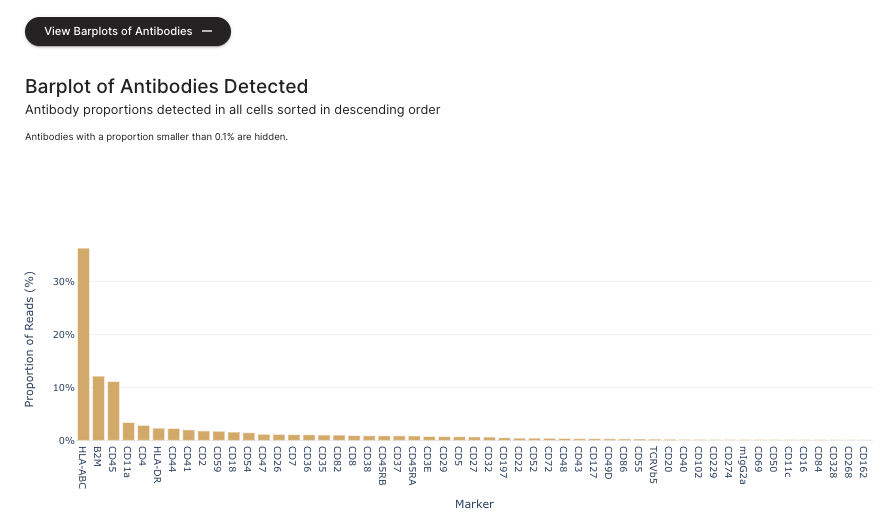
Sequencing
The sequencing section is showing different metrics that allows troubleshooting if parts of the amplicon sequenced have been subjected to any biases.
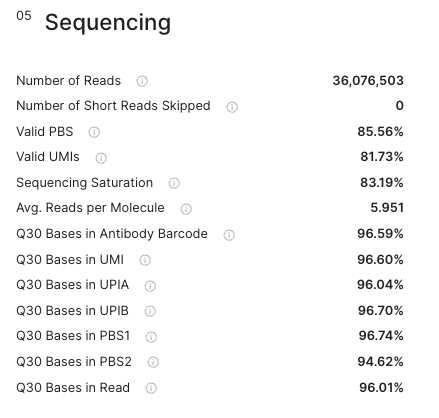
The dashboard on the right side displays a barplot that portrays the level of sequencing saturation of your MPX library. This can be toggled between absolute numbers and proportions for comparison purposes.
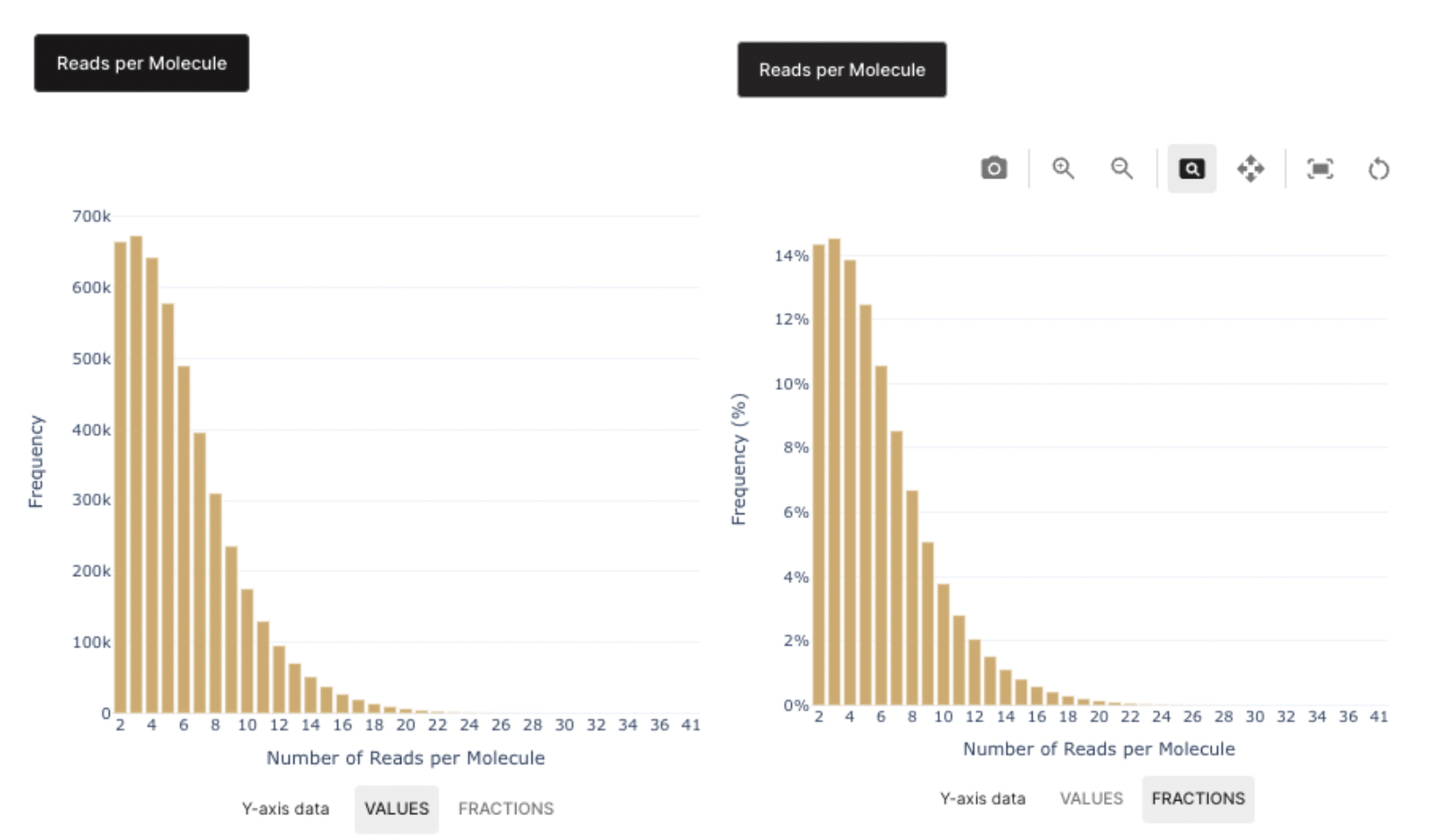
At the moment, pixelator doesn't compute a display of the saturation curve in the sequencing section.